Drug-target interaction is a prominent research area in drug discovery, which refers to the recognition of interactions between chemical compounds and the protein targets. Chemists estimate that 1060 compounds with drug-like properties could be made—that’s more than the total number of atoms in the Solar System, as an article reported in the journal Nature in 2017.
Drug development, on average, takes about 14 years and costs up to 1.5 billion dollars. During the journey of drug discovery in this vast “galaxy,” it is apparent that traditional biological experiments for DTI detection are normally costly and time-consuming.
Prof. Hou Tingjun is an expert in computer-aided drug design (CADD) at the Zhejiang University College of Pharmaceutical Sciences. In the past decades, he has been committed to developing drugs using computer technology. “The biggest challenge lies in the interactions between unknown targets and drug molecules. How can we discover them more efficiently? This involves a new breakthrough in method.”
Recently, artificial intelligence (AI) has opened up new possibilities. “With artificial intelligence, we may be able to reach the more upstream stage in drug discovery, thus improving the efficiency and success rate of the drug development,” said Hou.
In addition to AI, multi-omics data, such as genomics, proteomics, and pharmacology, have also flourished. In each field, there has been a massive ocean of biomedical information. The information about drugs, proteins, diseases, side effects, biological processes, molecular functions, cellular components, biological enzymes and ion channels has been storied in specialized databanks. However, their value for drug discovery remains obscure.
Prof. He Shibo is a scholar who specializes in big data and network science at the Zhejiang University College of Control Science and Engineering. “This domain is particularly suited for inter-disciplinary research. This considerable body of biological information can be abstracted into a multi-layered and heterogeneous network system,” said He.
In November 2021, Hou Tingjun, He Shibo and Cao Dongsheng at Central South University co-published a research article entitled “A unified drug-target interaction prediction framework based on knowledge graph and recommendation system” in the journal Nature Communications.
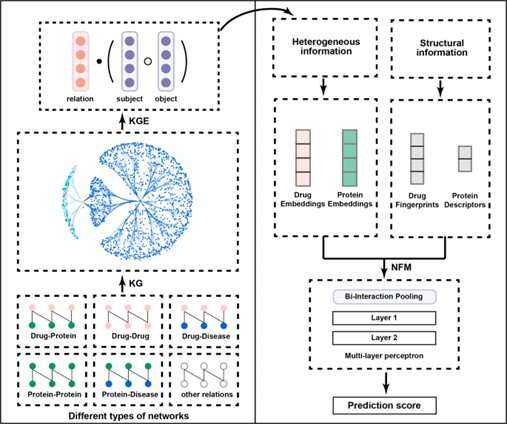
In this study, researchers proposed a unified framework called KGE_NFM (knowledge graph embedding and neural factorization machine) by incorporating KGE and recommendation system techniques for drug-target interactions (DTI) prediction that are applicable to the various scenarios of drug discovery, especially when encountering new protein targets.
Researchers evaluated KGE_NFM in three real-world scenarios: the warm start, the cold start for drugs and the cold start for proteins. In the first two scenarios, AI algorithms were on par with traditional ones, and sometimes even slightly inferior to the latter. In the third scenario, KGE_NFM outdistanced its rivals by 30%.
“This demonstrates the remarkable ability and superiority of AI in predicting the unknown protein targets. Discovering ‘the unknown drug-target interactions’ from ‘the unknown protein targets‘ is an undeniably important undertaking in the future of drug discovery,” Hou observed.
“We can do a lot of interesting things using AI for complex heterogeneous networking mining,” said He. For example, the team is currently working with a lab at Tencent to carry out research into virtual screening of hepatitis B drugs and drug synergy. “The use of KGE can not only expand the dimension of information but also promote the interpretability and credibility of algorithmic systems.”




















